It is generally believed that gene expression has a major impact on the development and effects of Alzheimer’s disease (AD) on the brain’s cognitive functions, but it is unclear how. Research depends on postmortem brain transcriptome profiles, which are the set of RNA transcripts produced by the genome. However, these profiles lack a temporal resolution, or in other words, it is impossible to determine how changes in these profiles occurred with time. To determine this relationship, researchers studied 30 AD-associated gene co-expression modules using fruit fly models. By studying the effects of amyloid beta (Aβ) plaques, tau neurofibrillary tangles, and aging, they were able to find 141 modifiers of induced neurodegeneration. Through genetic manipulation, there is an immune module enriched with AD variants (or alterations in the genetic sequence involved in the immune response to the disease), which then promotes neurodegeneration. Early AD activates expression of synaptic transcriptional signature (expressing a gene or genes related to the synapse) that promotes damage to neurons. In total, there is a causal link established from early AD, changes in the RNA transcripts, NMDA receptors, and neurodegeneration.
Microscopic of Neural network Brain cells
Image Source: fatido
As mentioned, a key challenge to studying the temporal relationship is exactly that: it is temporal in nature. The brain transcriptome profiles available for study are only those at death, which means the disease has already progressed to its end stages. However, the information needed is at the onset and evolution of the disease. This can be satisfied by using controlled animal models, where researchers can manipulate the genes and isolate the particular proteins similar to those in AD. Although initial attempts in mouse models worked well, analysis was limited between 500 and 5000 genes, which is where fruit flies come into play.
The specific genetic code used in transcriptome profiling is called UAS-tau, which exists in chromosome III and encodes a MAPT protein. Part of the code creates a 42-amino-acid peptide, while the other encodes alpha-synuclein (and had to be added to the genetic code of the fruit flies). This, plus crossing between flies, created the required genetic pool.
In previous analysis of postmortem brain RNA sequences found 3,774 AD differentially expressed genes and 30 brain gene co-expression network modules. Analysis based on Aβ and tau neuropathology, all 30 modules were confirmed to be associated with AD pathology. But as to which transcriptional changes were caused by AD and to differentiate between Aβ and tau proteins as the specific causes, we turn our attention to the fruit fly models. Expression of the 42-amino-acid peptide or the tau protein continues age-dependent dsyfunction and death of neurons and the central nervous system. Aβ induced 2,746 differentially expressed genes and tau induced 3,830 such genes (between 14-22% and 18-43% of genes in modules respectively), sharing the triggers for 1,448 genes. Mapping the data between humans and fruit flies, researchers found that most modules were induced by both proteins (with very low probability of these genes triggering by chance).
In human brains, these two proteins also appear alongside other proteins, such as alpha-synuclein, and this protein’s gene expression was also manipulated in these experiments (although note that alpha-synuclein is more commonly related to Parkinson’s than AD). It is also important to note that aging itself has an independent impact on the brain, and had the greatest impact, affecting over 10,000 genes. (αSyn affected the expression of 24-39% of genes in each human co-expression module, while aging affected between 86-95% of genes.)
From here, researchers developed a causal link in how triggers (Aβ, tau, and aging) affect brain gene expression networks, which then affect CNS function. To understand the connection between the brain transcriptional networks (the regulation of gene expression affecting neuron development and synaptic function) and CNS behavior, we add in locomotor performance. The genetic expression of 6 of 19 modules (the 19 modules varying similarly over all fly conditions) correlated heavily with locomotor performance, including networks involving genes for immune response, synaptic transmission, and endocytosis/vesicle trafficking. Aβ triggered increased expression of the immune response, which then affected the locomotor impairment caused by Aβ. On the other hand, the tau protein caused decreased expression for synaptic transmission, and only partially affected locomotor impairment.
One module triggered by the proteins and/or aging is the STGblue module, as its gene expression increases and can be transcribed for more AD variants. The second module affected is the PHGbrown module, which regulates synaptic transmission. This module contains genes that encode suppressors of Aβ and tau, and becomes less expressed. Without regulation of the synapses, neurons may become overexcited or die completely from poor development. Overall, the three aforementioned triggers, along with αSyn, are responsible for roughly 86% of changes in gene expression within tested modules, while the remaining could be explained by other pathologies causing dementia and other brain diseases.
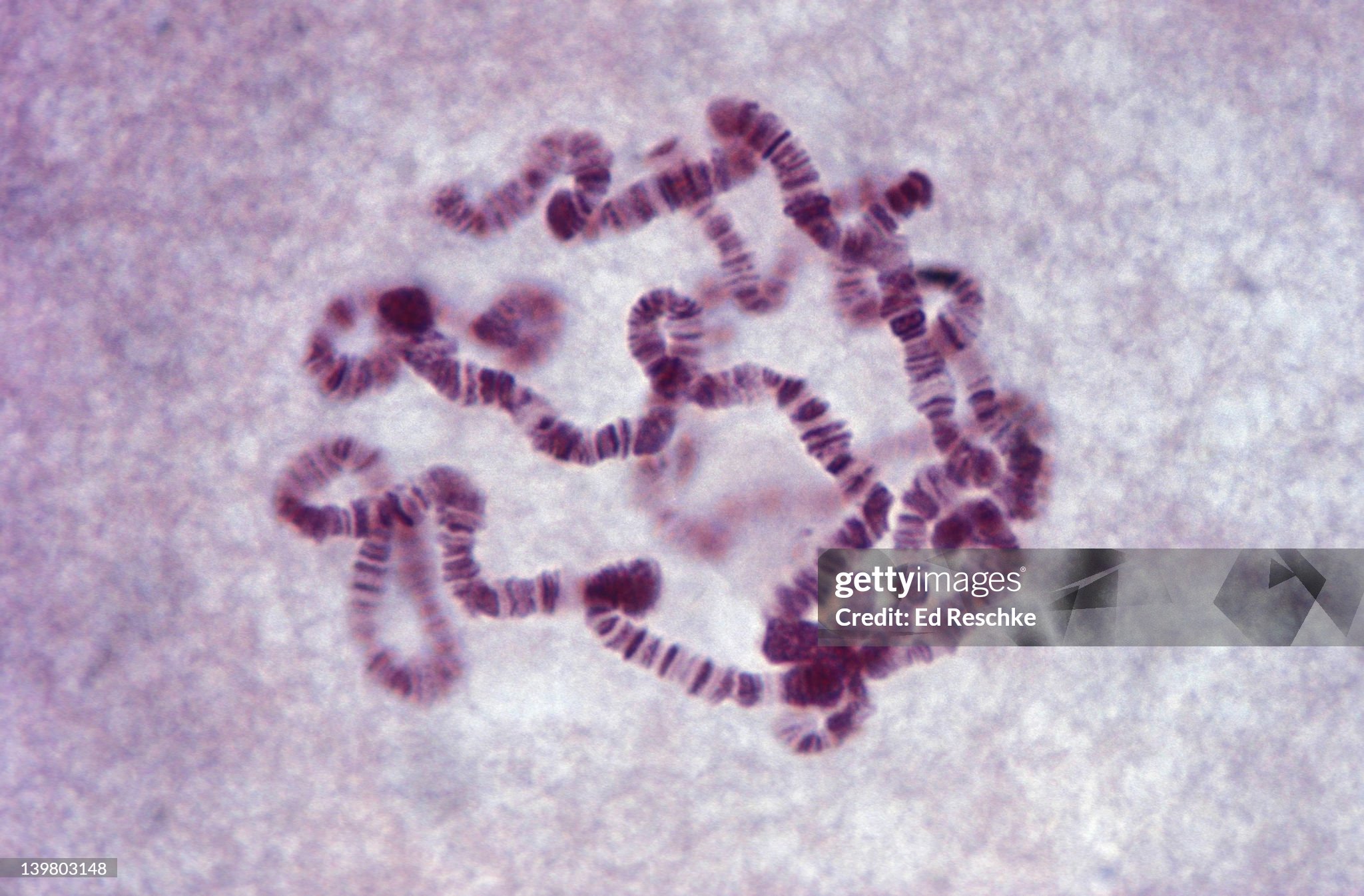
Giant Chromosomes, Drosophila (Fruitfly) from Salivary Glands of Larva Shows Bands. 280X
Image Source: Ed_Reschke
Featured Image Source: ShinneProject